This is an AI for Health MSc project. Students are eligible to receive a monthly reimbursement of €500,- for a period of six months. For more information please read the requirements.
Clinical problem
Cerebral Small Vessel Disease (CSVD) causes a fifth of all stroke cases and is the major vascular cause of cognitive decline and dementia. CSVD is a neurological disorder present in virtually every individual aged 60 years old or older consisting of pathological processes that disturb cerebral arterioles, capillaries and venules, leading to white and grey matter lesions. As it is not possible to visualize the smallest vasculature in vivo yet, we focus on understanding the underlying pathology of the radiological hallmarks linked to CSVD (e.g. White matter hyperintensities (WMH)).
WMH , seen as hyperintense on FLAIR- and T2-weighted MRI sequences, are amongst the foremost recurrent CSVD features observed on MRI scans and represent different degrees of axonal loss, demyelination and gliosis. Despite the fact that WMH hold a strong association with an increased risk of long-term cognitive impairment, little is known about their underlying pathology. Studies focusing on pathological examination on WMH until now have shown semi-quantitative quantification methodologies, which might lead to subjective data analysis, and to mischaracterization of the pathological processes behind these lesions. Diffusion Tensor Imaging (DTI) has shown that WMH are surrounded by penumbra regions with different diffusion metrics that appear normal on conventional MRI. These penumbra regions are believed to closely correlate to WMH progression. However, imaging-pathology studies to this date have failed to capture the earliest changes in WMH, and thereby of those mechanisms preceding their progression.
Therefore, there is a need for an automatic analysis of WMH progression across imaging, and its detailed correlation to pathological markers. We aim to research these through a postmortem human study that combines high field (7T) MRI with immuno-histochemical (IHC) stainings on sections of the same brain regions. To this date, few studies have examined the relationship between radiological and pathological correlates in CSVD, however none of them has provided a systematic and detailed description of the pathology behind WMH. Our dataset provides the unique opportunity to accurately characterize pathological changes of multiple neurodegenerative processes in grey matter (GM), normal appearing white matter (NAWM) and WMH.
Solution
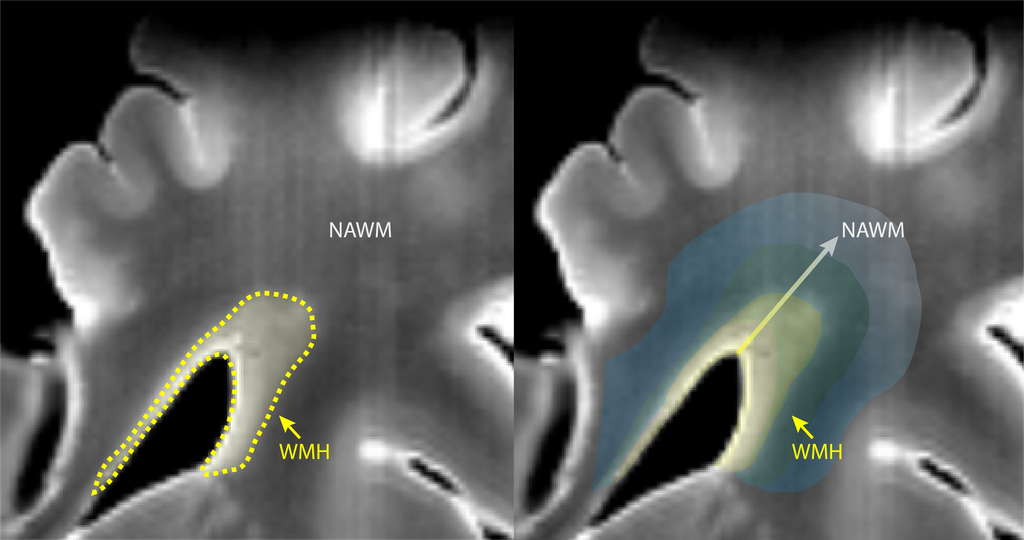
Based on previous studies combining histopathological examination together with MRI, it is possible to identify and characterize several CSVD-linked lesions. Thus far, we have realigned both MRI imaging and IHC sections, and we have performed manual segmentation of grey matter (GM), normal appearing white matter (NAWM) and WMH (Figure 1). However, this is not enough to study the transition from the core of the lesion to healthy white matter tissue. We believe that by characterizing the pathological changes occurring across white matter from lesion towards healthy tissue we will obtain a unique insight into the key processes leading to WMH expansion. In order to do so, we would like to combine our manual segmentation data together with automatic segmentation of the gradual changes in WMH towards the so-called NAWM. We believe that by including this analysis on our dataset we might be able to gain a unique insight into changes occurring within white matter tissue beyond current knowledge and how these could be characterized on standard MRI, or currently available alternatives e.g. DTI.
Within this AI project we would like to characterize WMH in detail as well as transition zones that are visible in MRI (via an automatic segmentation pipeline). Specifically, we will use an existing white matter lesion segmentation network developed at the Radboudumc on the inv vivo MRIs over 400 patients (the RUN-DMC dataset). We will adapt this network to ex vivo MRI data, which is mostly similar in appearance. Then apply these identified subregions in the ex vivo MRI are then mapped to our different histopathological markers, and thereby further the understanding of white matter changes that have been overlooked until now because of methodological limitation.
Data
In order to answer this question, we have a data set with HF MRI and histopathology of 28 individuals (approximately 5 TB). Regarding MRI, we acquired T1, T2, T2* sequences at 400 x 400 x 400 µm resolution and FLAIR at 500 x 500 x 500 µm. To study WMH we focused on T2-w MRI, which was used to assess WMH burden by volumetric segmentation. WMH can be divided into periventricular WMH (PVWMH) and deep WMH (DWMH). PVWMH has been proven to correlate to cognitive decline, therefore we performed biopsies of the tissue adjacent to the left lateral ventricle. Most of these individuals were sampled twice, so approximately 60 biopsies were made. Each of these biopsies corresponds to a 2.5 x 2 x 2 cm tissue slabs that were further sectioned at 4 µm. Several stainings have been performed such as Hematoxylin/Eosin (HE), Luxol Fast Blue (LFB) and with antibodies against Neurofilament (NF), Glial Fibrillary Acidic Protein (GFAP), Amyloid-ß (Aß) and microglia activation (Ionized calcium binding adaptor molecule 1, Iba-1). Additionally, we stained for vascular integrity via Glucose Transporter 1 (GLUT1), Smooth muscle 1 (SM1) and elastin through Masson’s Trichrome staining (MTS). All stainings were performed at the Pathology department from the Radboud University Medical Center, Nijmegen, The Netherlands. In addition, we have an existing deep learning model developed on in vivo MRIs of 400 patients as a starting point for model training and development. For this project in particular , the student will mainly focus on the analysis of T2-w MRI data. 2D imaging-pathology co-registration has already been performed and WMH have been manually segmented. However, this fails to recapitulate the progression of WMH. Therefore, the student will focus on classification of white matter into different clusters based on the burden observed on MRI.
Results
An automatic classifier to assess WMH burden beyond the standard division of lesion or “healthy” brain tissue, will provide novel clinical meaningful insights in the characterization of CSVD and its prognosis. WMH are known to correlate to cognitive impairment. Studies researching the progression of WMH have found that this is not always linear, and that higher burden of WMH hold the strongest association with cognitive decline. However, current imaging approaches fail to predict how these lesions will progress as analysis tends to focus solely on WMH. Higher field strength MRI and DTI can entail more details than currently considered for the evaluation of these lesions. The automatic analysis tools applied within this project can be used to obtain unbiased results within larger clinical datasets and other research projects. A fully automatic analysis pipeline overcomes interrater differences as well as facilitates the unification of several study results to improve the understanding of a mechanism and/or disease. Furthermore, this postmortem study on CSVD is in combination with the departments of Neurology and Geriatric Medicine, hoping that our discoveries can aid in the regular clinical practice too. The information from our current study, is closely linked to the research within the department of Neurology, where there are several studies focusing on understanding CSVD progression (RUN DMC study). Finally, by investigating changes in white matter and how these might differently impact pathology depending on the burden of WMH, we expect to elucidate which mechanisms play a role in later stages of disease, and ultimately lead to dementia. Altogether, we believe that the possibility of developing an automatic classifier for white matter changes from our HF MRI dataset can be beneficial for research, and to clinical practice from speeding up diagnostic procedures to help understand the progression of white matter lesions.
Embedding
You will be embedded in the Department Medical Imaging, Anatomy at Radboudumc, and collaborate closely with the Computational Pathology Group Radboud UMC. We provide access to several analysis PCs, direct supervision by PhD students and regular meetings with the supervisors of this project.
Requirements
- Students Artificial Intelligence, Data Science, Computer Science, Bioinformatics, or Biomedical Sciences in the final stages of their Master education.
- You should be proficient in python programming and have a theoretical understanding of deep learning
- Basic biological / biomedical knowledge is preferred.
Information
- Project duration: 6 months
- Location: Radboud University Medical Center
- For more information or to apply for this project, please contact Geert.Litjens@radboudumc.nl.
